15 results
Characterization of carbapenem-resistant gram-negative bacteria collected in the Sentinel Surveillance Program, 2018–2019
- Lori Spicer, Davina Campbell, J. Kristie Johnson, Cynthia Longo, Thomas Balbuena, Thomas Ewing, Maria Karlsson, J. Kamile Rasheed, Christopher Elkins, Amy Gargis, Joseph Lutgring
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, p. s52
-
- Article
-
- You have access Access
- Open access
- Export citation
-
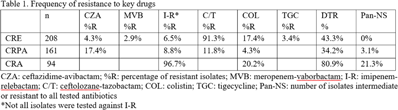
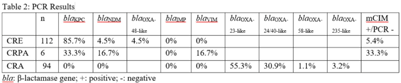
Background: Carbapenem resistance in gram-negative organisms is an important public health problem. The CDC conducted Sentinel surveillance in 2018–2019 to characterize these organisms from 9 facilities in 9 different states. Methods: Carbapenem-resistant Enterobacterales (CRE), Pseudomonas aeruginosa (CRPA), and Acinetobacter spp (CRA) obtained from clinical samples of patients in acute-care or long-term care facilities were submitted to the CDC. Identification was confirmed using matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF), and antimicrobial susceptibility testing (AST) was performed via broth microdilution for 27 antibiotics. All confirmed CRE and CRPA were tested for carbapenemase production (CP) using the modified carbapenem inactivation method (mCIM). The isolates that were mCIM-positive were assessed by real-time PCR for presence of blaKPC, blaNDM, blaVIM, and blaIMP. CP-CRE were also assessed for blaOXA-48-like. All confirmed CRA were tested for the same genes as CRPA and blaOXA-23–like, blaOXA-24/40-like, blaOXA-58–like, and blaOXA-235–like genes. Difficult-to-treat resistance (DTR) was defined as resistance to all β-lactams (excluding newer β-lactam combination agents) and quinolones tested. Results: The CDC confirmed 208 CRE, 161 CRPA, and 94 CRA. Table 1 summarizes AST results for a selection of drugs. We identified 112 (53.8%) mCIM-positive CRE and 6 (3.7%) mCIM-positive CRPA. The PCR results are summarized in Table 2. One mCIM-positive and PCR-negative isolate was positive in a metallo-β-lactamase screen. Conclusions: Resistance among CRE and CRPA to newer β-lactam combination agents was detected. Options for treating CRA are limited. Of 112 CP-CRE, 85.7% harbored blaKPC; CP-CRPA were rare (3.7%); and most CRA harbored blaOXA-23-like (55.3%) or blaOXA-24/40-like (30.9%). Whole-genome sequencing is planned to better understand gene variants, sequence types, and additional resistance markers present among the isolates.
Funding: None
Disclosures: None
Epidemiology of extended-spectrum β-lactamase–producing Enterobacterales in five US sites participating in the Emerging Infections Program, 2017
- Nadezhda Duffy, Maria Karlsson, Hannah E. Reses, Davina Campbell, Jonathan Daniels, Richard A. Stanton, Sarah J. Janelle, Kyle Schutz, Wendy Bamberg, Paulina A. Rebolledo, Chris Bower, Rebekah Blakney, Jesse T. Jacob, Erin C. Phipps, Kristina G. Flores, Ghinwa Dumyati, Hannah Kopin, Rebecca Tsay, Marion A. Kainer, Daniel Muleta, Benji Byrd-Warner, Julian E. Grass, Joseph D. Lutgring, J. Kamile Rasheed, Christopher A. Elkins, Shelley S. Magill, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 43 / Issue 11 / November 2022
- Published online by Cambridge University Press:
- 14 February 2022, pp. 1586-1594
- Print publication:
- November 2022
-
- Article
- Export citation
-
Objective
The incidence of infections from extended-spectrum β-lactamase (ESBL)–producing Enterobacterales (ESBL-E) is increasing in the United States. We describe the epidemiology of ESBL-E at 5 Emerging Infections Program (EIP) sites.
MethodsDuring October–December 2017, we piloted active laboratory- and population-based (New York, New Mexico, Tennessee) or sentinel (Colorado, Georgia) ESBL-E surveillance. An incident case was the first isolation from normally sterile body sites or urine of Escherichia coli or Klebsiella pneumoniae/oxytoca resistant to ≥1 extended-spectrum cephalosporin and nonresistant to all carbapenems tested at a clinical laboratory from a surveillance area resident in a 30-day period. Demographic and clinical data were obtained from medical records. The Centers for Disease Control and Prevention (CDC) performed reference antimicrobial susceptibility testing and whole-genome sequencing on a convenience sample of case isolates.
ResultsWe identified 884 incident cases. The estimated annual incidence in sites conducting population-based surveillance was 199.7 per 100,000 population. Overall, 800 isolates (96%) were from urine, and 790 (89%) were E. coli. Also, 393 cases (47%) were community-associated. Among 136 isolates (15%) tested at the CDC, 122 (90%) met the surveillance definition phenotype; 114 (93%) of 122 were shown to be ESBL producers by clavulanate testing. In total, 111 (97%) of confirmed ESBL producers harbored a blaCTX-M gene. Among ESBL-producing E. coli isolates, 52 (54%) were ST131; 44% of these cases were community associated.
ConclusionsThe burden of ESBL-E was high across surveillance sites, with nearly half of cases acquired in the community. EIP has implemented ongoing ESBL-E surveillance to inform prevention efforts, particularly in the community and to watch for the emergence of new ESBL-E strains.
Carbapenem-Resistant Enterobacteriaceae Resistant Only to Ertapenem: An Epidemiologically Distinct Cohort, Atlanta, 2016–2018
- Chris Bower, Max Adelman, Jessica Howard-Anderson, Uzma Ansari, Joseph Lutgring, Gebre Tiga, Jesse Jacob
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s463-s464
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenem-resistant Enterobacteriaceae (CRE), particularly carbapenemase-producing (CP) CRE, pose a major public health threat. In 2016, the phenotypic definition of CRE expanded to include ertapenem resistance to improve sensitivity for detecting CP-CRE. We compared characteristics of CRE resistant to ertapenem only (CRE-EO) to CRE resistant to ≥1 other carbapenem (CRE-O). Methods: The Georgia Emerging Infections Program performs active, population-based CRE surveillance in metropolitan Atlanta. CRE cases were defined as any Escherichia coli, Klebsiella pneumoniae, K. oxytoca, K. variicola, Enterobacter cloacae complex, or Enterobacter aerogenes resistant to ≥1 carbapenem by the clinical laboratory and isolated from urine or a sterile site between 2016 and 2018. Data were extracted from retrospective chart review and 90-day mortality from Georgia vital statistics for 2016–2017. Polymerase chain reaction (PCR) for carbapenemase genes was performed on a convenience sample of isolates by the CDC or Georgia Public Health Laboratory. We compared characteristics of CRE-EO cases to CRE-O cases using χ2 tests or t tests. Results: Among 927 CRE isolates, 553 (60%) were CRE-EO. CRE-EO were less frequently isolated from blood (5% vs 12%; P < .01) and less commonly K. pneumoniae (21% vs 58%; P < .01) than CRE-O. CRE-EO cases were more often women (65% vs 50%; P < .01), had a lower Charlson comorbidity index (mean ± SD, 2.4±2.3 vs 3.0±2.6; P < .01), and were less commonly at a long-term care facility (24% vs 31%) or hospital (15% vs 21%; P < .01) in the 4 days prior to the CRE culture. CRE-EO were more susceptible to all antibiotics tested at the clinical laboratory (P < .01) except for tigecycline (P = 1.0) (Table 1). Of the 300 (32%) isolates tested for carbapenemase genes, 98 (33%) were positive (7% CRE-EO vs 62% CRE-O; P < .01). Of the CP isolates, we identified blaKPC in 93 cases (95%), blaNDM in 3 cases (3%), blaOXA-48-like in 2 cases (2%). CRE-EO cases had lower 90-day mortality (13% vs 21%; P < .01). Conclusions: CRE-EO are epidemiologically distinct from CRE-O and are less likely to harbor carbapenemase genes. CRE-EO may require less intensive infection prevention interventions and have more therapeutic options.
Funding: None
Disclosures: None
Trimethoprim-Sulfamethoxazole Resistance Patterns Among Methicillin-Resistant Staphylococcus aureus, 2012–2018
- D. Cal Ham, Lucy Fike, Tara Fulton, Nychie Dotson, Rebecca Perlmutter, Ericka Kalp, Joseph Lutgring, Amy Gargis, Alison Laufer-Halpin, Alexander Kallen, Maroya Walters
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s418
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Trimethoprim-sulfamethoxazole is commonly used for the treatment of noninvasive methicillin-resistant Staphylococcus aureus (MRSA) infections. Following a report from 2 facilities of increased trimethoprim-sulfamethoxazole resistance among MRSA infections, we assessed changes in resistance nationally and by state. Methods: We reviewed antibiotic susceptibility testing (AST) data for trimethoprim-sulfamethoxazole among S. aureus isolates associated with surgical site infections (SSIs), central-line–associated bloodstream infections (CLABSIs), and catheter-associated urinary tract infections (CAUTIs) from acute-care hospitals reported to the NHSN Device and Procedure Module from 2012 to 2018. We compared the pooled mean percentage of isolates nonsusceptible to trimethoprim-sulfamethoxazole in 2012 and 2018, stratified by MRSA and methicillin-sensitive Staphylococcus aureus (MSSA). Among MRSA isolates, we compared the percentage nonsusceptible to trimethoprim-sulfamethoxazole by healthcare-associated infection (HAI) type and state in 2012 and 2018. States with ≥20 MRSA isolates with AST reported each year were included in the state-level analysis. Results: Overall, 36,587 MRSA isolates and 46,824 MSSA isolates were reported from 2012 to 2018. Moreover, >80% of MRSA and MSSA isolates had trimethoprim-sulfamethoxazole AST reported each year. Nationally, the percentage of trimethoprim-sulfamethoxazole nonsusceptible among MRSA isolates was 3.9% in 2012 compared to 6.5% in 2018 (P < .001), but it was unchanged among MSSA isolates during the same period (1.1% in 2012 vs 1.4% in 2018; P = .08). Among MRSA surgical site infections (SSIs), the proportion of trimethoprim-sulfamethoxazole nonsusceptible isolates was 3.1% in 2012 versus 6.1% in 2018 (P < .001) but did not change significantly for CLABSIs or CAUTIs (Fig. 1). Among the 32 states that met the inclusion criteria, there were no significant decreases, whereas 4 (12.5%) showed significant increases in the percentage of MRSA that were trimethoprim-sulfamethoxazole nonsusceptible in 2018 compared to 2012: New Jersey (2.4% in 2012 vs 19.3% in 2018; P <.001); Florida (9.1% in 2012 vs 22.4% in 2018; P < .001); Maryland (0.0% in 2012 vs 10.9% in 2018; P < .01); and Pennsylvania (1.7% in 2012 vs 6.5% in 2018; P < .001). Conclusions: Nationally, there was a modest but significant increase in the percentage of MRSA HAI isolates nonsusceptible to trimethoprim-sulfamethoxazole in 2018 compared to 2012; however, 3 of 4 states with significant increases in nonsusceptibility had substantial, potentially clinically relevant increases (>10%). Ongoing characterization of MRSA isolates from Florida and New Jersey may provide insight into the underlying cause of these shifting patterns in trimethoprim-sulfamethoxazole resistance among MRSA. Healthcare personnel should select appropriate antibiotic regimens based on local resistance patterns, should monitor patients for treatment failure, and should report changes in resistance to the appropriate public health department.
Funding: None
Disclosures: None
Evaluation of Discrepancies in Carbapenem Minimum Inhibitory Concentrations Obtained at Clinical Laboratories Compared to a Public Health Laboratory
- Julian E. Grass, Shelley S. Magill, Isaac See, Uzma Ansari, Lucy E. Wilson, Elisabeth Vaeth, Paula Snippes Vagnone, Brittany Pattee, Jesse T. Jacob, Georgia Emerging Infections Program, Chris Bower, Atlanta Veterans Affairs Medical Center, Foundation for Atlanta Veterans Education and Research, Sarah W. Satola, Sarah J. Janelle, Kyle Schutz, Rebecca Tsay, Marion A. Kainer, Daniel Muleta, P. Maureen Cassidy, Vivian H. Leung, Meghan Maloney, Erin C. Phipps, New Mexico Emerging Infections Program, Kristina G. Flores, New Mexico Emerging Infections Program, Erin Epson, Joelle Nadle, Maria Karlsson, Joseph D. Lutgring
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s474-s476
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Automated testing instruments (ATIs) are commonly used by clinical microbiology laboratories to perform antimicrobial susceptibility testing (AST), whereas public health laboratories may use established reference methods such as broth microdilution (BMD). We investigated discrepancies in carbapenem minimum inhibitory concentrations (MICs) among Enterobacteriaceae tested by clinical laboratory ATIs and by reference BMD at the CDC. Methods: During 2016–2018, we conducted laboratory- and population-based surveillance for carbapenem-resistant Enterobacteriaceae (CRE) through the CDC Emerging Infections Program (EIP) sites (10 sites by 2018). We defined an incident case as the first isolation of Enterobacter spp (E. cloacae complex or E. aerogenes), Escherichia coli, Klebsiella pneumoniae, K. oxytoca, or K. variicola resistant to doripenem, ertapenem, imipenem, or meropenem from normally sterile sites or urine identified from a resident of the EIP catchment area in a 30-day period. Cases had isolates that were determined to be carbapenem-resistant by clinical laboratory ATI MICs (MicroScan, BD Phoenix, or VITEK 2) or by other methods, using current Clinical and Laboratory Standards Institute (CLSI) criteria. A convenience sample of these isolates was tested by reference BMD at the CDC according to CLSI guidelines. Results: Overall, 1,787 isolates from 112 clinical laboratories were tested by BMD at the CDC. Of these, clinical laboratory ATI MIC results were available for 1,638 (91.7%); 855 (52.2%) from 71 clinical laboratories did not confirm as CRE at the CDC. Nonconfirming isolates were tested on either a MicroScan (235 of 462; 50.9%), BD Phoenix (249 of 411; 60.6%), or VITEK 2 (371 of 765; 48.5%). Lack of confirmation was most common among E. coli (62.2% of E. coli isolates tested) and Enterobacter spp (61.4% of Enterobacter isolates tested) (Fig. 1A), and among isolates testing resistant to ertapenem by the clinical laboratory ATI (52.1%, Fig. 1B). Of the 1,388 isolates resistant to ertapenem in the clinical laboratory, 1,006 (72.5%) were resistant only to ertapenem. Of the 855 nonconfirming isolates, 638 (74.6%) were resistant only to ertapenem based on clinical laboratory ATI MICs. Conclusions: Nonconfirming isolates were widespread across laboratories and ATIs. Lack of confirmation was most common among E. coli and Enterobacter spp. Among nonconfirming isolates, most were resistant only to ertapenem. These findings may suggest that ATIs overcall resistance to ertapenem or that isolate transport and storage conditions affect ertapenem resistance. Further investigation into this lack of confirmation is needed, and CRE case identification in public health surveillance may need to account for this phenomenon.
Funding: None
Disclosures: None
Characterization of Ceftazidime-Avibactam-Resistant Carbapenem-Resistant Enterobacteriaceae, United States, 2015–2017
- Uzma Ansari, Hannah E. Reses, Julian Grass, Joelle Nadle, Chris Bower, Jesse Jacob, Elisabeth Vaeth, Medora Witwer, Emily Hancock, Suzanne Dale, Ghinwa Dumyati, Zintars Beldavs, Daniel Muleta, Nadezhda Duffy, Isaac See, Joseph Lutgring
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s465-s466
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenem-resistant Enterobacteriaceae (CRE) are a major public health problem. Ceftazidime-avibactam (CZA) is a treatment option for CRE approved in 2015; however, it does not have activity against isolates with metallo-β-lactamases (MBLs). Emerging resistance to CZA is a cause for concern. Our objective was to describe the microbiologic and epidemiologic characteristics of CZA-resistant (CZA-R) CRE. Methods: From 2015 to 2017, 9 states participated in laboratory- and population-based surveillance for carbapenem-resistant Escherichia coli, Klebsiella pneumoniae, K. oxytoca, K. aerogenes, and Enterobacter cloacae complex isolates from a normally sterile site or urine. A convenience sample of isolates from this surveillance were sent to the CDC for antimicrobial susceptibility testing (AST) using reference broth microdilution (BMD) including an MBL screen, species confirmation with MALDI-TOF, and real-time PCR to detect blaKPC, blaNDM, and blaOXA-48–like genes. Additional AST by BMD was performed on CZA-R isolates using meropenem-vaborbactam (MEV), imipenem-relebactam (IMR), plazomicin (PLZ), and eravacycline (ERV). Epidemiologic data were obtained from a medical record review. Community-associated cases were defined as having no healthcare exposures in the year prior to culture, no devices in place 2 days prior to culture, and culture collected before calendar day 3 after hospital admission. Data were analyzed in 3 groups: CRE that were CZA-susceptible (CZA-S), CZA-R that were due to blaNDM, and CZA-R without blaNDM. Results: Among 606 confirmed CRE tested with CZA, 33 (5.4%) were CZA-R. Of the CZA-R isolates, 16 (48.5%) harbored a blaNDM gene, of which 2 coharbored blaNDM and blaOXA-48-like genes; 9 (27.3%) harbored only a blaKPC gene. Of the 17 CZA-R isolates without blaNDM, all were MBL screen negative. CZA-R due to blaNDM were more frequently community-associated (43.8%) than CZA-S or CZA-R without blaNDM (11.0% and 5.9%, respectively); a higher percentage of CZA-R cases due to blaNDM also had recent international travel (25%) compared to the other groups (1.8% and 5.9%, respectively). CZA-R without blaNDM were more susceptible to MEV (76%), IMR (71%), PLZ (88%), and ERV (65%) compared to CZA-R due to blaNDM (19%, 6%, 56%, and 44%, respectively). Conclusions: The emergence of CZA-R isolates without blaNDM are concerning; however, these isolates are more susceptible to newer antimicrobials than those with blaNDM. In addition to high rates of resistance to newer antimicrobials, isolates with blaNDM are more frequently community-associated than other CRE. This underscores the need for more aggressive measures to stop the spread of CRE.
Funding: None
Disclosures: None
Pilot Program for Aztreonam-Avibactam Susceptibility Testing of Metallo-Beta-Lactamase-Producing Enterobacteriaceae
- Amelia Bhatnagar, Sarah Malik, Maria Karlsson, David Lonsway, Joseph Lutgring, Jennifer Huang, Stephanie Gumbis, Allison Brown
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s74-s75
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenemase-producing Enterobacteriaceae (CPE) are a major public health concern because they typically display multidrug resistance and they cause hard-to-treat infections. Organisms harboring metallo-β-lactamases (MBLs) pose a critical challenge in clinical practice because they confer resistance to nearly all β-lactams, including recently approved β-lactam combination agents. A promising new β-lactam-β-lactamase inhibitor combination for treating infections caused by MBL-producing CPE is aztreonam–avibactam. Although clinical trials using aztreonam–avibactam are ongoing, clinicians can administer this combination using 2 US Food and Drug Administration (FDA)–approved drugs: aztreonam and ceftazidime–avibactam. In 2019, the Centers for Disease Control and Prevention (CDC) initiated a pilot program in the Antibiotic Resistance Laboratory Network (AR Lab Network) to address the lack of commercially available antimicrobial susceptibility tests (ASTs) for aztreonam-avibactam by performing broth microdilution (BMD) for this drug combination. We describe the isolates submitted for aztreonam-avibactam AST during the AR Lab Network pilot in 2019. Methods: The AR Lab Network regional laboratories adopted the HP D300e Digital Dispenser to create customized BMD panels for aztreonam–avibactam ASTs. To qualify for aztreonam–avibactam AST, isolates had to be an Enterobacteriaceae displaying nonsusceptibility to all tested β-lactams (including either ceftazidime-avibactam or meropenem-vaborbactam) or confirmed to harbor at least 1 MBL gene (blaVIM, blaNDM, or blaIMP). Regional laboratories confirmed carbapenemase gene(s) using a molecular method. If an MBL gene was confirmed, aztreonam-–avibactam minimum inhibitory concentrations (MICs) were reported back to submitters within 3 working days of receipt. Findings were reported to CDC using a REDCap database. Results: From March through August 2019, aztreonam–avibactam AST was requested for 32 clinical isolates across 16 states. These isolates included 15 Escherichia coli, 12 Klebsiella pneumoniae, 4 Enterobacter cloacae complex, and 1 Proteus mirabilis. Molecular detection identified 27 blaNDM-positive isolates, 2 blaOXA-48-like-positive isolates, and 3 blaOXA-48/blaNDM-positive isolates. Aztreonam-avibactam results were reported for 30 isolates; 5 displayed elevated aztreonam-avibactam MICs of 8/4 µg/mL (n = 4) or 16/4 µg/mL (n = 1). Results for 2 isolates were not reported because the isolates were MBL negative. Aztreonam-avibactam MICs ranged from 0.06/4 µg/mL to 16/4 µg/mL. The MIC50/MIC90 were 0.5/4 µg/mL and 8/4 µg/mL. Conclusions: In the absence of effective FDA-approved treatments and lack of available AST for novel antibiotic combinations, CDC’s provision of AST for aztreonam-avibactam among MBL-producing CPE, offered through the AR Lab Network, helps fill a critical gap to inform patient treatment decisions. To date, our in vitro data suggest that aztreonam–avibactam could be a promising drug combination for use against infections caused by MBL-producing Enterobacteriaceae.
Funding: None
Disclosures: None
Molecular Typing of Invasive Staphylococcus aureus from the Emerging Infections Program (EIP) Using Whole-Genome Sequencing
- Davina Campbell, Gillian McAllister, Kelly Jackson, Isaac See, Alison Halpin, Joseph Lutgring, Erin Epson, Susan Petit, Susan Ray, William Schaffner, Ghinwa Dumyati, Thomas Ewing, Michelle Adamczyk, Amy Gargis
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s71-s72
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: The CDC has performed surveillance for invasive Staphylococcus aureus (iSA) infections through the Emerging Infections Program (EIP) since 2004. SCCmec and spa typing for clonal complex (CC) assignment and genomic markers have been used to characterize isolates. In 2019, whole-genome sequencing (WGS) of isolates began, allowing for high-resolution assessment of genomic diversity. Here, we evaluate the reliability of SCCmec typing, spa typing, and CC assignment using WGS data compared to traditional methods to ensure that backwards compatibility is maintained. Methods:S. aureus isolates were obtained from a convenience sample of iSA cases reported through the EIP surveillance system. Overall, 78 iSA isolates with diverse spa repeat patterns, CCs, SCCmec types, and antimicrobial susceptibility profiles were sequenced (MiSeq, Illumina). Real-time PCR and Sanger sequencing were used as the SCCmec and spa typing reference methods, respectively. spa-MLST mapping (Ridom SpaServer) served as the reference method for CC assignment. WGS assembly and multilocus sequence typing (MLST) were performed using the CDC QuAISAR-H pipeline. WGS-based MLST CCs were assigned using eBURST and SCCmec types using SCCmecFinder. spa types were assigned from WGS assemblies using BioNumerics. For isolate subtyping, previously published and validated canonical single-nucleotide polymorphisms (canSNPs) as well as the presence of the Panton-Valentine leukocidin (PVL) toxin and arginine catabolic mobile element (ACME) virulence factor were assessed for all genome assemblies. Results: All isolates were assigned WGS-based spa types, which were 100% concordant (78 of 78) with Sanger-based spa typing. SCCmecFinder assigned 91% of isolates (71 of 78) SCCmec types, which were 100% concordant with reference method results. Also, 7 isolates had multiple cassettes predicted or an incomplete SCCmec region assembly. Using WGS data, 96% (75 of 78) of isolates were assigned CCs; 3 isolates had unknown sequence types that were single-locus variants of established sequence types. Overall, 70 isolates had CCs assigned by the reference method; 100% (70 of 70) concordance was observed with WGS-based CCs. Analysis of canSNPs placed 42% (33 of 78) of isolates into CC8, with 17 (52%) of these isolates classified as USA300. PVL and ACME were not accurate markers for inferring the USA300 subtype as 24% (4 of 17) of isolates did not contain these markers. Conclusions:S. aureus CCs, SCCmec, and spa types can be reliably determined using WGS. Incorporation of canSNP analysis represents a more efficient method for CC8 assignment than the use of genomic markers alone. WGS allows for the replacement of multiple typing methods for increased laboratory efficiency, while maintaining backward compatibility with historical typing nomenclature.
Funding: None
Disclosures: None
Whole-Genome Sequencing Reveals Diversity of Carbapenem-Resistant Pseudomonas aeruginosa Collected Through the Emerging Infections Program
- Richard Stanton, Jonathan Daniels, Erin Breaker, Davina Campbell, Joseph Lutgring, Maria Karlsson, Kyle Schutz, Jesse Jacob, Lucy Wilson, Elisabeth Vaeth, Linda Li, Ruth Lynfield, Erin C. Phipps, Emily Hancock, Ghinwa Dumyati, Rebecca Tsay, P. Maureen Cassidy, Jacquelyn Mounsey, Julian Grass, Maroya Walters, Alison Halpin
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s513-s514
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenem-resistant Pseudomonas aeruginosa (CRPA) is a frequent cause of healthcare-associated infections (HAIs). The CDC Emerging Infections Program (EIP) conducted population and laboratory-based surveillance of CRPA in selected areas in 8 states from August 1, 2016, through July 31, 2018. We aimed to describe the molecular epidemiology and mechanisms of resistance of CRPA isolates collected through this surveillance. Methods: We defined a case as the first isolate of P. aeruginosa resistant to imipenem, meropenem, or doripenem from the lower respiratory tract, urine, wounds, or normally sterile sites identified from a resident of the EIP catchment area in a 30-day period; EIP sites submitted a systematic random sample of isolates to CDC for further characterization. Of 1,021 CRPA clinical isolates submitted, 707 have been sequenced to date using an Illumina MiSeq. Sequenced genomes were classified using the 7-gene multilocus sequence typing (MLST) scheme, and a core genome MLST (cgMLST) scheme was used to determine phylogeny. Antimicrobial resistance genes were identified using publicly available databases, and chromosomal mechanisms of carbapenem resistance were determined using previously validated genetic markers. Results: There were 189 sequence types (STs) among the 707 sequenced genomes (Fig. 1). The most frequently occurring were high-risk clones ST235 (8.5%) and ST298 (4.7%), which were found across all EIP sites. Carbapenemase genes were identified in 5 (<1%) isolates. Overall, 95.6% of the isolates had chromosomal mutations associated with carbapenem resistance: 93.2% had porinD-associated mutations that decrease membrane permeability to the drugs; 24.8% had mutations associated with overexpression of the multidrug efflux pump MexAB-OprM; and 22.9% had mutations associated with overexpression of the endogenous β-lactamase ampC. More than 1 such chromosomal resistance mutation type was present in 37.8% of the isolates. Conclusions: The diversity of the sequence types demonstrates that HAIs caused by CRPA can arise from a variety of strains and that high-risk clones are broadly disseminated across the EIP sites but are a minority of CRPA strains overall. Carbapenem resistance in P. aeruginosa was predominantly driven by chromosomal mutations rather than acquired mechanisms (ie, carbapenemases). The diversity of the CRPA isolates and the lack of carbapenemase genes suggest that this ubiquitous pathogen can readily evolve chromosomal resistance mechanisms, but unlike carbapenemases, these cannot be easily spread through horizontal transfer.
Funding: None
Disclosures: None
Molecular Epidemiology and Outcomes of Patients with Carbapenem-Resistant Enterobacteriaceae Bacteriuria, Atlanta 2012–2015
- Jessica Howard-Anderson, Robert Petit, Chris Bower, Gillian Smith, Uzma Ansari, Alison Halpin, Maria Karlsson, Adrian Lawson, Joseph Lutgring, Gillian McAllister, Monica Farley, Jesse Jacob, Sarah Satola
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s489-s490
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Carbapenem-resistant Enterobacteriaceae (CRE) represent a significant antibiotic resistance threat, in part because carbapenemase genes can spread on mobile genetic elements. Here, we describe the molecular epidemiology and outcomes of patients with CRE bacteriuria from the same city in a nonoutbreak setting. Methods: The Georgia Emerging Infections Program performs active, population-based CRE surveillance in Atlanta. We studied a cohort of patients with CRE (resistant to all tested third-generation cephalosporins and ≥1 carbapenem, excluding ertapenem) first identified in urine, and not in a prior or simultaneous sterile site, between 2012 and 2015. Whole-genome sequencing (WGS) was performed on a convenience sample. We obtained epidemiologic and outcome data through chart review and Georgia Vital Statistics records (90-day mortality). Using WGS, we created a core-genome alignment-based phylogenetic tree of the Klebsiella pneumoniae isolates and calculated the SNP difference between each sample. Using SAS version 9.4 software, we performed the Fisher exact test and univariable odds ratios (OR) with 95% CI to compare patient isolates with and without a carbapenemase gene. Results: Among 81 patients included, the median age was 68 (IQR, 57–74) years, and most were female (58%), black (60%), and resided in a long-term care facility 4 days prior to culture isolation (53%). Organisms isolated were K. pneumoniae (84%), Escherichia coli (7%), Enterobacter cloacae (7%), and Klebsiella oxytoca (1%). WGS identified at least 1 β-lactamase gene in 91% of the isolates; 85% contained a carbapenemase gene, the most frequent of which was blaKPC-3 (94%). Patients with CRE containing a carbapenemase gene were more likely to be black (OR, 3.7; 95% CI, 1.0–13.8) and to have K. pneumoniae (OR, 8.9; 95% CI, 2.2–35.0). Using a core-genome alignment of 3,708 genes (~63% of the complete genome), we identified a median of 67 (IQR, 23–3,881) SNP differences between each K. pneumoniae isolate. A phylogenetic tree identified clustering around carbapenemase gene and multilocus sequence type (84% were ST 258) but not based on referring laboratory or county of residence (Fig. 1). Although 7% of patients developed an invasive CRE infection within 1 year and 21% died within 90 days, having a carbapenemase gene was not associated with these outcomes. Conclusions: Molecular sequencing of a convenience sample of CRE bacteriuria support K. pneumoniae ST258 harboring blaKPC-3 being distributed throughout the Atlanta area, across the healthcare continuum. Overall mortality was high in this population, but the presence of carbapenemase genes was not associated with worse outcomes.
Funding: None
Disclosures: None
Disclosures: None
Funding: None
Chlorhexidine MICs Remain Stable Among Antibiotic-Resistant Bacterial Isolates Collected from 2005 to 2019 at Three US Sites
- Joseph Lutgring, Julian Grass, David Lonsway, Brian Yoo, Erin Epson, Megan Crumpler, Karen Galliher, Matthew Zahn, Eric Evans, Jesse Jacob, Alexander Page, Sarah Satola, Gillian Smith, Marion Kainer, Mary Hayden, Sujan Reddy, Christopher Elkins, Shelley Magill, Alice Guh
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s26
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Chlorhexidine bathing reduces bacterial skin colonization and prevents infections in specific patient populations. As chlorhexidine use becomes more widespread, concerns about bacterial tolerance to chlorhexidine have increased; however, testing for chlorhexidine minimum inhibitory concentrations (MICs) is challenging. We adapted a broth microdilution (BMD) method to determine whether chlorhexidine MICs changed over time among 4 important healthcare-associated pathogens. Methods: Antibiotic-resistant bacterial isolates (Staphylococcus aureus from 2005 to 2019 and Escherichia coli, Klebsiella pneumoniae, and Enterobacter cloacae complex from 2011 to 2019) were collected through Emerging Infections Program surveillance in 2 sites (Georgia and Tennessee) or through public health reporting in 1 site (Orange County, California). A convenience sample of isolates were collected from facilities with varying amounts of chlorhexidine use. We performed BMD testing using laboratory-developed panels with chlorhexidine digluconate concentrations ranging from 0.125 to 64 μg/mL. After successfully establishing reproducibility with quality control organisms, 3 laboratories performed MIC testing. For each organism, epidemiological cutoff values (ECVs) were established using ECOFFinder. Results: Among 538 isolates tested (129 S. aureus, 158 E. coli, 142 K. pneumoniae, and 109 E. cloacae complex), S. aureus, E. coli, K. pneumoniae, and E. cloacae complex ECVs were 8, 4, 64, and 64 µg/mL, respectively (Table 1). Moreover, 14 isolates had an MIC above the ECV (12 E. coli and 2 E. cloacae complex). The MIC50 of each species is reported over time (Table 2). Conclusions: Using an adapted BMD method, we found that chlorhexidine MICs did not increase over time among a limited sample of S. aureus, E. coli, K. pneumoniae, and E. cloacae complex isolates. Although these results are reassuring, continued surveillance for elevated chlorhexidine MICs in isolates from patients with well-characterized chlorhexidine exposure is needed as chlorhexidine use increases.
Funding: None
Disclosures: None
Epidemiologic Characteristics of ESBL-Producing ST131 E. coli Identified Through the Emerging Infections Program, 2017
- Nadezhda Duffy, Davina Campbell, Hannah E. Reses, Elizabeth Basiliere, Chris Bower, Ghinwa Dumyati, Marion Kainer, Daniel Muleta, Benji Byrd-Warner, Richard Stanton, Jonathan Daniels, Alison Halpin, Joseph Lutgring, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s214-s215
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Extended-spectrum β-lactamase–producing (ESBL) Escherichia coli infection incidence is increasing in the United States. This increase may be due to the rapid expansion of ST131, which is now the predominant ESBL strain globally, often multidrug resistant, and has been shown to establish longer-term human colonization than other E. coli strains. We assessed potential risk factors that distinguish ST131 from other ESBL E. coli. Methods: From October 1 through December 31, 2017, 5 CDC Emerging Infections Program (EIP) sites pilot tested active, laboratory-based surveillance in selected counties in Colorado, Georgia, New Mexico, New York, and Tennessee. An E. coli case was defined as the first isolation from a normally sterile body site or urine in a surveillance area resident in a 30-day period resistant to 1 extended-spectrum cephalosporin antibiotic and susceptible or intermediate to all carbapenem antibiotics tested. Epidemiologic data were collected from case patients’ medical records. A convenience sample of 117 E. coli isolates from case patients was collected. All isolates underwent whole-genome sequencing to determine sequence type and the presence of ESBL genes. We compared ST131 E. coli epidemiology to other ESBL E. coli. Results: Among 117 E. coli isolates, 97 (83%) were ESBL producers. Of the 97 ESBL E. coli, 52 (54%) were ST131 (range, for 4 EIP sites submitting >10 isolates: 25%–88%; P < .001). Other common STs were ST38 (12%) and ST10 (5%). ST131 infections were more likely to be healthcare-associated than non-ST131 (56% vs 36%; P = .05) (Table 1). Among specific prior healthcare exposures, only residence in long-term care facilities (LTCFs) in the year before culture was more common among ST131 case patients (29% vs 11%; P = .03). Notably, 85% of ESBL E. coli collected from LTCF residents were ST131. ST131 E. coli were more common among patients with underlying medical conditions (81% vs 60%; P = .02). No statistically significant difference by sex, race, age, culture source, location of culture collection, and frequency of antibiotic use in the prior 30 days was observed. Conclusions:The prevalence of ST131 E. coli varies regionally. The association between ST131 and LTCFs suggests that these may be particularly important settings for ST131 acquisition. Improving infection control measures that limit ESBL transmission in these settings and preventing dissemination in facilities receiving patients from LTCFs may be necessary to contain ST131 spread.
Funding: None
Disclosures: None
Antimicrobial Nonsusceptibility Among Invasive MRSA USA300 Strains by Healthcare Exposure, Three Sites, 2005–2016
- Kelly Jackson, Runa Gokhale, Davina Campbell, Amy Gargis, Susan Ray, Ruth Lynfield, William Schaffner, Joseph Lutgring, Isaac See
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s120-s121
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Incidence of community-associated (CA) and healthcare-associated, community-onset (HACO) USA300 methicillin-resistant Staphylococcus aureus (MRSA) bloodstream infections has remained unchanged in recent years. Traditionally considered a CA strain, USA300 is increasingly associated with healthcare settings. We examined whether antimicrobial nonsusceptibility among USA300 strains could distinguish epidemiologic class (community vs hospital), and whether divergences in susceptibility were occurring over time. Methods: We used data on invasive MRSA infections from active, population, and laboratory-based surveillance during 2005–2016 from 11 counties in 3 states. Invasive cases were defined as MRSA isolation from a normally sterile site in a surveillance area resident. Cases were considered hospital-onset (HO) if the culture was obtained >3 days after hospitalization and HACO if ≥1 of the following risk factors was present: hospitalization, surgery, dialysis, or residence in a long-term care facility in the past year; or central vascular catheter ≤2 days before culture. Otherwise, cases were considered CA. Sites submitted a convenience sample of clinical MRSA isolates for molecular typing and antimicrobial susceptibility testing. Molecular typing was performed by pulsed-field gel electrophoresis until 2008, when typing was inferred using a validated algorithm based on molecular characteristics. Reference broth microdilution was performed for 8 antimicrobials and interpreted based on CLSI interpretive criteria. We compared USA300 nonsusceptibility for HO and CA isolates. For antimicrobials with >5% nonsusceptibility and for which HO isolates had greater nonsusceptibility than CA isolates, we compared nonsusceptibility for HACO and CA and analyzed annual trends in nonsusceptibility within each epidemiologic class (ie, CA, HACO, and HO) using linear regression. Results: Of 17,947 MRSA cases during 2005–2016, isolates were available for 6,685 (37%), and 2,120 were USA300 (34% CA, 52% HACO, 14% HO). HO isolates had more nonsusceptibility than CA isolates to gentamicin (2.2% vs 0.6%; P = .03), levofloxacin (47.8% vs 39.7%; P = .02), rifampin (3.7 vs 1.1%; P = .01), and trimethoprim-sulfamethoxazole (3.4% vs 0.6%; P = .04). HACO isolates also had more nonsusceptibility than CA isolates to levofloxacin (50.9% vs 39.7%; P < .01). Levofloxacin nonsusceptibility increased during 2005–2016 for HACO and CA isolates (P < .01), but not among HO isolates (P = .36) (Fig. 1). Conclusions: Overall, nonsusceptibility across drugs cannot distinguish USA300 isolates causing HO versus CA disease. Although HO isolates had higher levofloxacin nonsusceptibility than CA and HACO isolates early on, USA300 MRSA HACO isolates now have levofloxacin nonsusceptibility most similar to that of HO isolates. Further study could help to explore whether increases in fluoroquinolone nonsusceptibility among CA and HACO cases may be contributing to the persistence of USA300 strains.
Disclosures: None
Funding: None
Multiple importations and transmission of colistin-resistant Klebsiella pneumoniae in a hospital in northern India
- Purva Mathur, Surbhi Khurana, Tom J.B. de Man, Neha Rastogi, Omika Katoch, Balaji Veeraraghavan, Ayyan Raj Neeravi, Manigandan Venkatesan, Subodh Kumar, Sushma Sagar, Amit Gupta, Richa Aggarwal, Kapil Dev Soni, Rajesh Malhotra, Anoop Velayudhan, Valan Siromany, Paul Malpiedi, Joseph Lutgring, Kayla Laserson, Neil Gupta, Padmini Srikantiah, Aditya Sharma
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 40 / Issue 12 / December 2019
- Published online by Cambridge University Press:
- 18 October 2019, pp. 1387-1393
- Print publication:
- December 2019
-
- Article
- Export citation
-
Objective:
Resistance to colistin, a last resort antibiotic, has emerged in India. We investigated colistin-resistant Klebsiella pneumoniae(ColR-KP) in a hospital in India to describe infections, characterize resistance of isolates, compare concordance of detection methods, and identify transmission events.
Design:Retrospective observational study.
Methods:Case-patients were defined as individuals from whom ColR-KP was isolated from a clinical specimen between January 2016 and October 2017. Isolates resistant to colistin by Vitek 2 were confirmed by broth microdilution (BMD). Isolates underwent colistin susceptibility testing by disk diffusion and whole-genome sequencing. Medical records were reviewed.
Results:Of 846 K. pneumoniae isolates, 34 (4%) were colistin resistant. In total, 22 case-patients were identified. Most (90%) were male; their median age was 33 years. Half were transferred from another hospital; 45% died. Case-patients were admitted for a median of 14 days before detection of ColR-KP. Also, 7 case-patients (32%) received colistin before detection of ColR-KP. All isolates were resistant to carbapenems and susceptible to tigecycline. Isolates resistant to colistin by Vitek 2 were also resistant by BMD; 2 ColR-KP isolates were resistant by disk diffusion. Moreover, 8 multilocus sequence types were identified. Isolates were negative for mobile colistin resistance (mcr) genes. Based on sequencing analysis, in-hospital transmission may have occurred with 8 case-patients (38%).
Conclusions:Multiple infections caused by highly resistant, mcr-negative ColR-KP with substantial mortality were identified. Disk diffusion correlated poorly with Vitek 2 and BMD for detection of ColR-KP. Sequencing indicated multiple importation and in-hospital transmission events. Enhanced detection for ColR-KP may be warranted in India.
Fecal Microbiota Transplant for Multidrug-Resistant Organism Decolonization Administered During Septic Shock
- Srinivasa Nithin Gopalsamy, Amy Sherman, Michael H. Woodworth, Joseph D. Lutgring, Colleen S. Kraft
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 39 / Issue 4 / April 2018
- Published online by Cambridge University Press:
- 18 January 2018, pp. 490-492
- Print publication:
- April 2018
-
- Article
-
- You have access Access
- HTML
- Export citation





